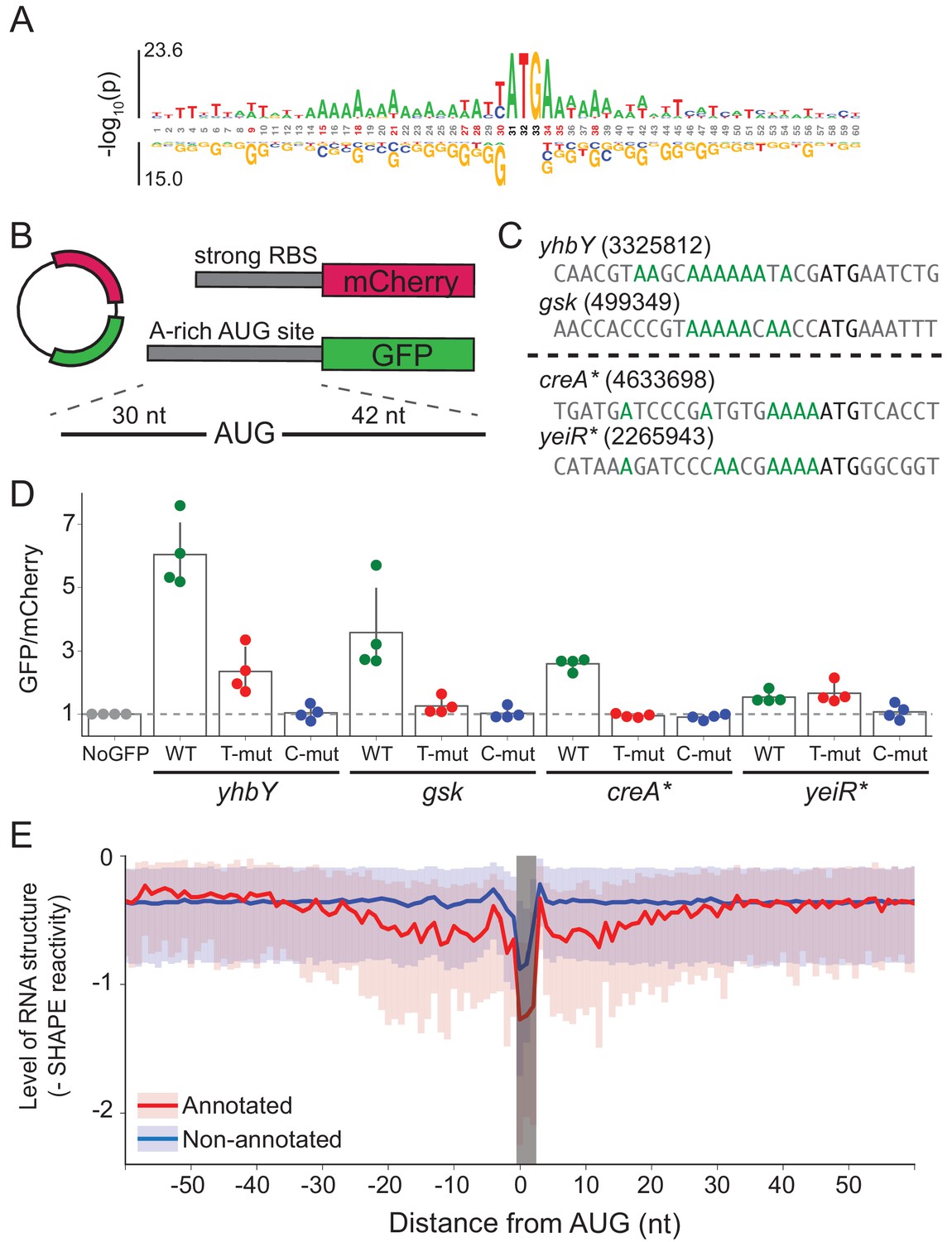
Translational initiation in E. coli occurs at the correct sites genome-wide in the absence of mRNA-rRNA base-pairing | eLife

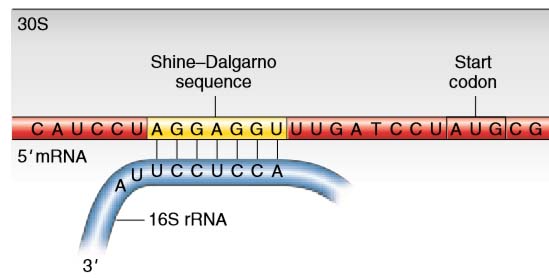
Twitter 上的 BASE mRNA Facility:"We were all taught the Shine-Dalgarno sequence but did you know it was discovered in Australia? The SD sequence is a ribosomal binding site in bacterial #mRNA that
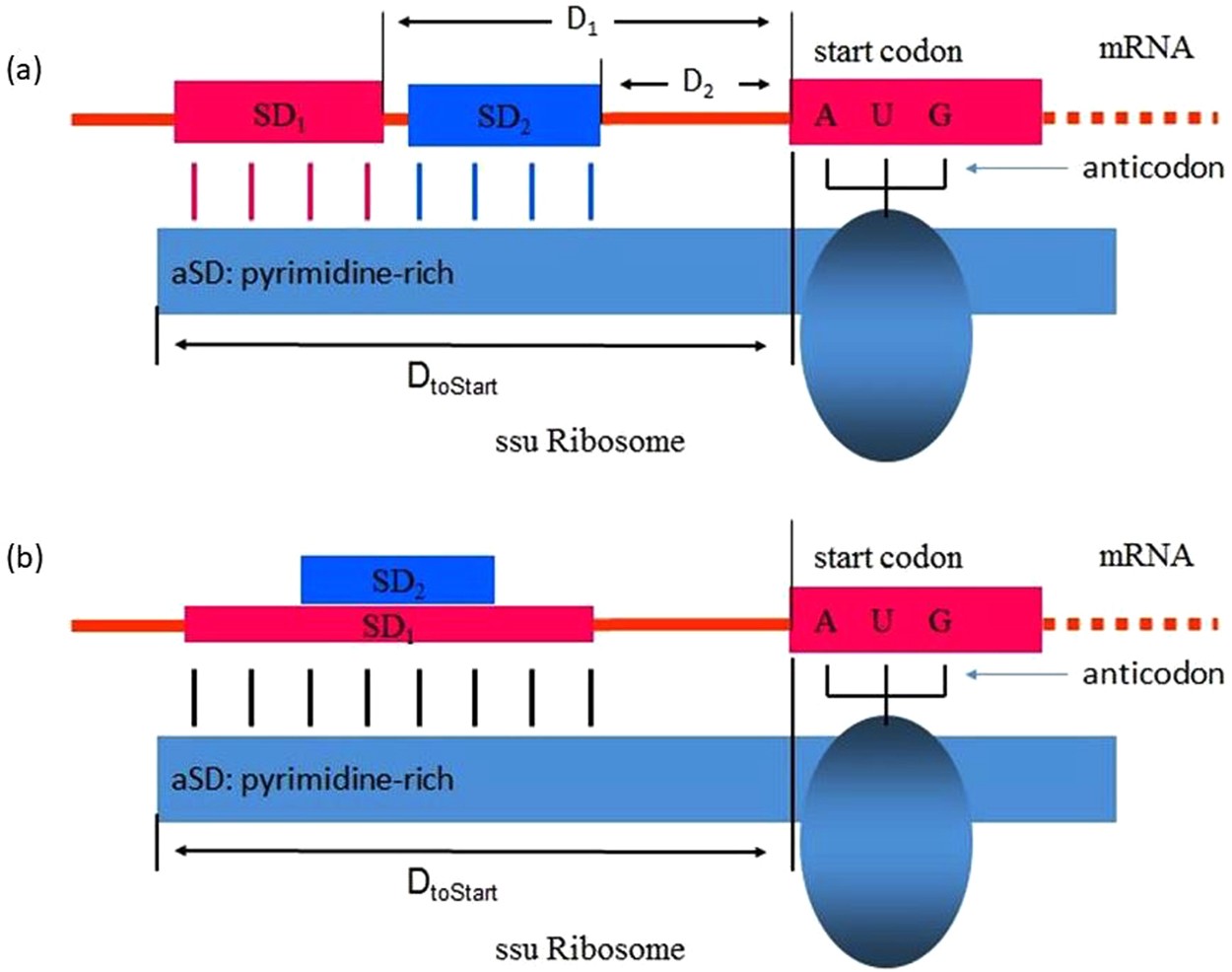
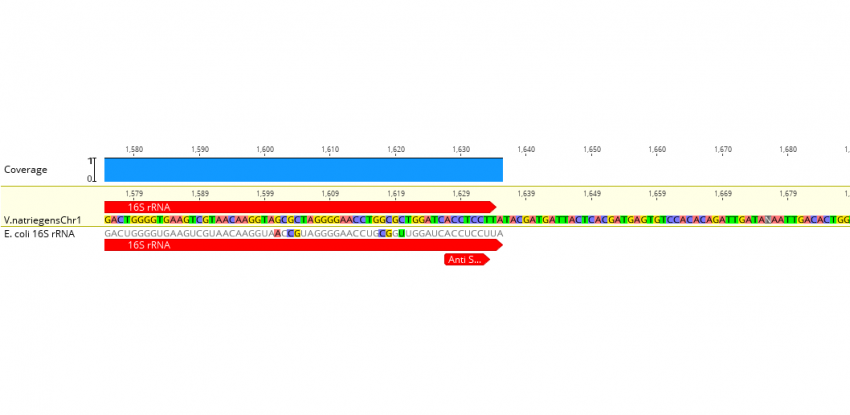
Elucidating the 16S rRNA 3′ boundaries and defining optimal SD/aSD pairing in Escherichia coli and Bacillus subtilis using RNA-Seq data | Scientific Reports

The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts | Nature Communications

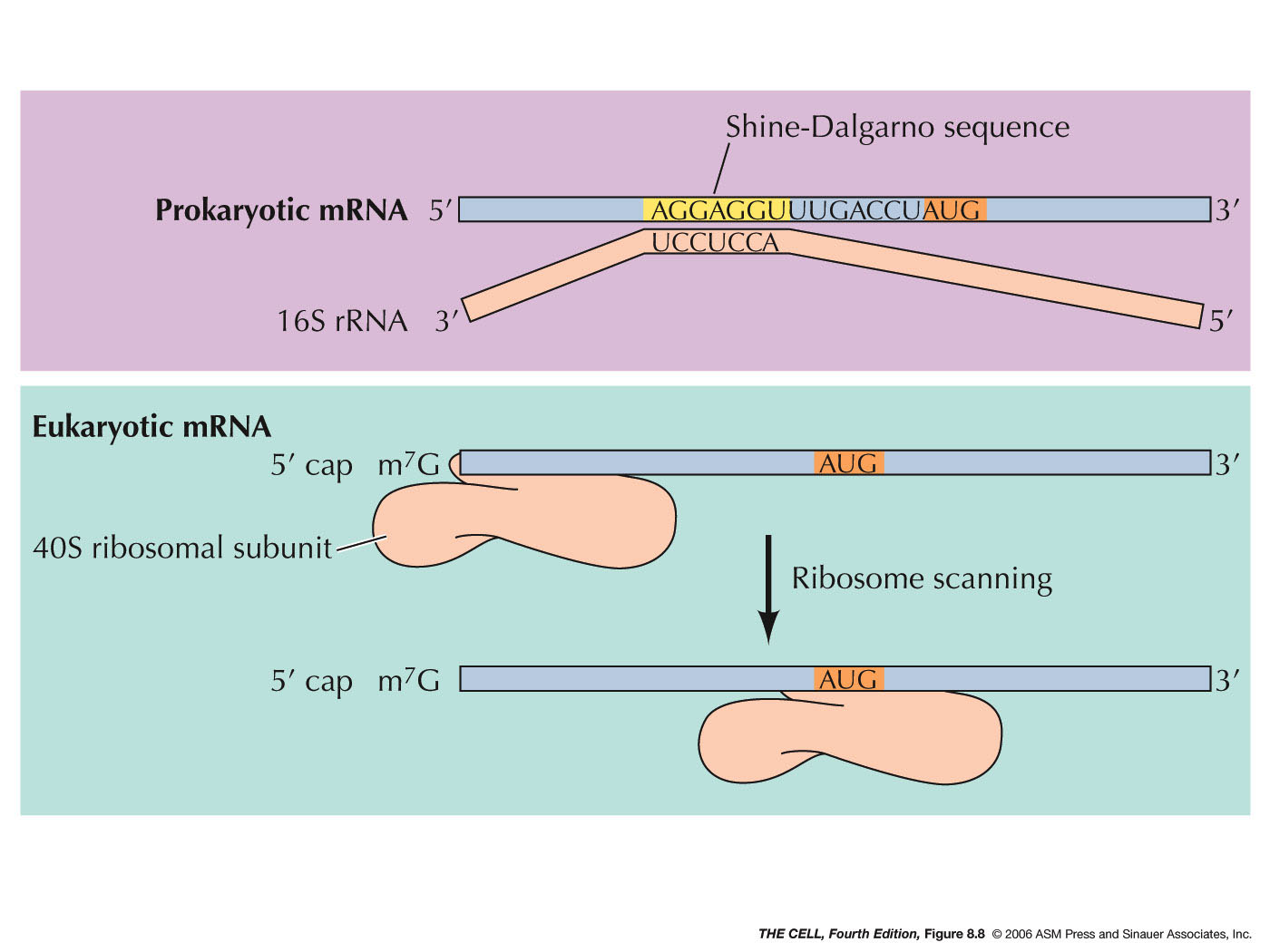
The possible dual impacts of Shine-Dalgarno (SD) sequences on protein... | Download Scientific Diagram

Leveraging genome-wide datasets to quantify the functional role of the anti- Shine–Dalgarno sequence in regulating translation efficiency | Open Biology