In silico analysis of the protein sequence of MNS1. BLAST analysis of... | Download Scientific Diagram

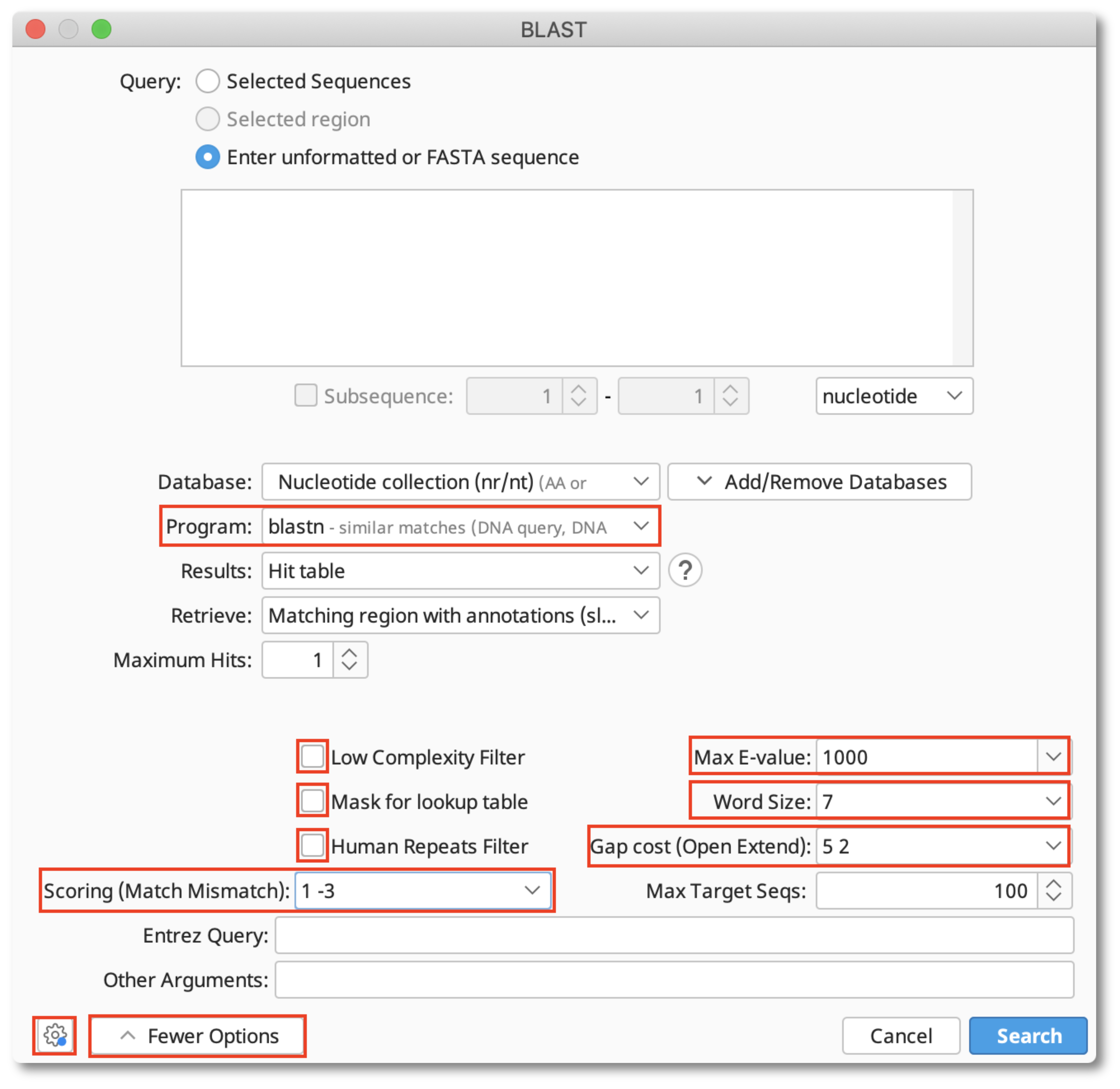
BLAST: Compare & identify sequences - NCBI Bioinformatics Resources: An Introduction - Library Guides at UC Berkeley

BLASTP Protein Sequence Search - Introduction to NCBI Bioinformatics Resources - Learning Resource Center at Uniformed Services University of the Health Sciences
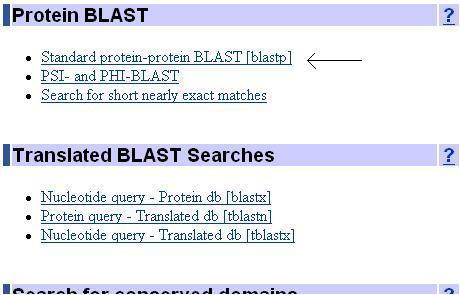
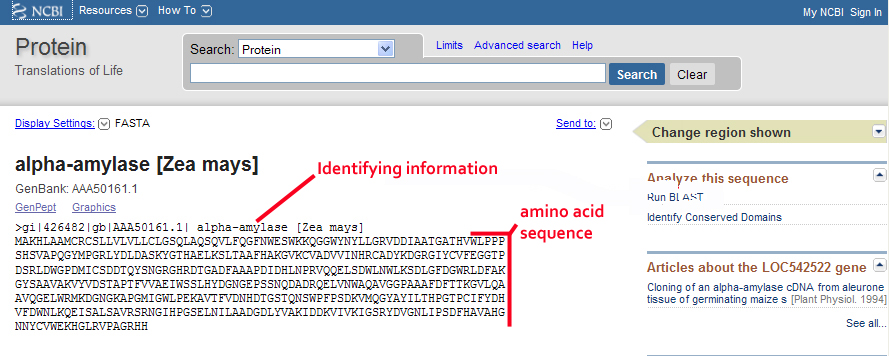
Protein BLAST allows one to input protein sequences and compare these against other protein sequences

BLAST: Compare & identify sequences - NCBI Bioinformatics Resources: An Introduction - Library Guides at UC Berkeley

BLAST Results - Introduction to NCBI Bioinformatics Resources - Learning Resource Center at Uniformed Services University of the Health Sciences

What is BLAST? BLAST® (Basic Local Alignment Search Tool) is a set of similarity search programs designed to explore all of the available sequence databases. - ppt video online download

The protein BLAST (BLASTp) search against the annotated NCBI protein... | Download Scientific Diagram
A) BLAST results of 10 AMP sequences with the novel designed peptide.... | Download Scientific Diagram




![Help [PIR - Protein Information Resource] Help [PIR - Protein Information Resource]](https://proteininformationresource.org/pirwww/images/creationpirsf.jpg)