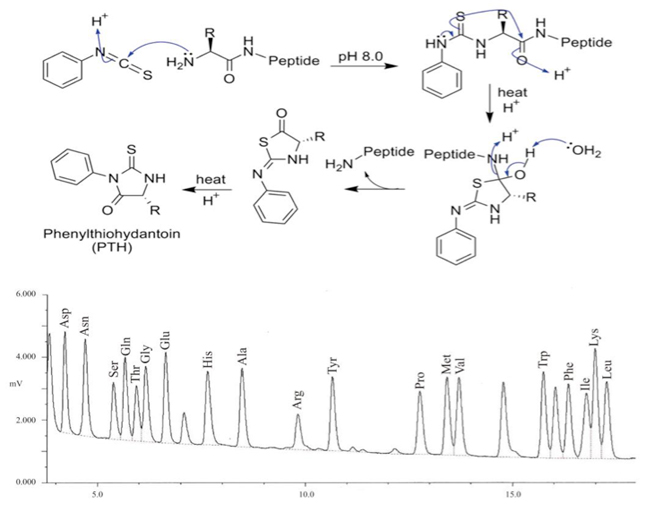
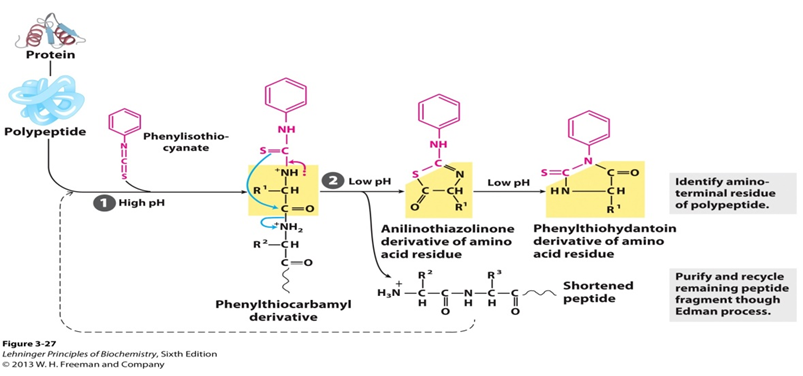
![Combining [13C6]‐phenylisothiocyanate and the Edman degradation reaction: a possible breakthrough for absolute quantitative proteomics together with protein identification - Oe - 2010 - Rapid Communications in Mass Spectrometry - Wiley Online Library Combining [13C6]‐phenylisothiocyanate and the Edman degradation reaction: a possible breakthrough for absolute quantitative proteomics together with protein identification - Oe - 2010 - Rapid Communications in Mass Spectrometry - Wiley Online Library](https://analyticalsciencejournals.onlinelibrary.wiley.com/cms/asset/342bd0c3-1a96-43b3-a3cb-641f85082d15/mfig001.jpg)
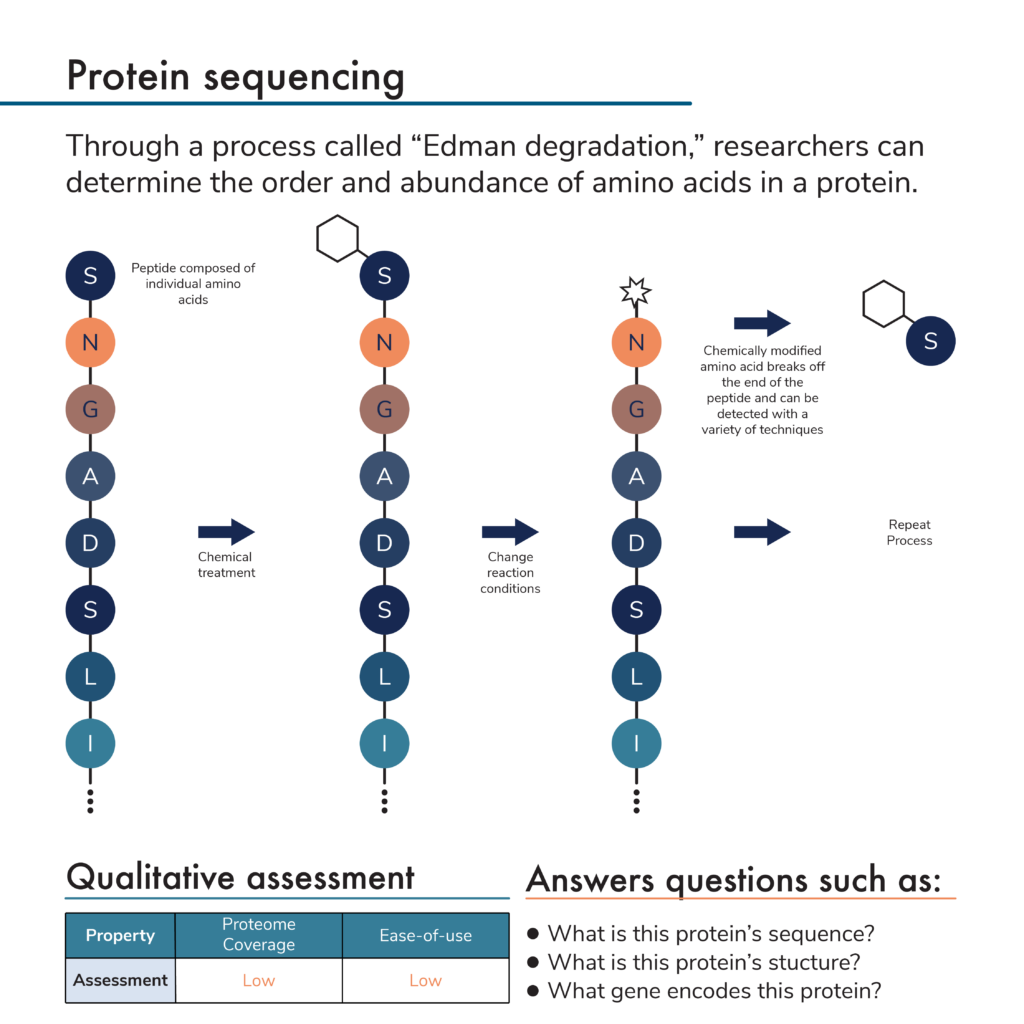
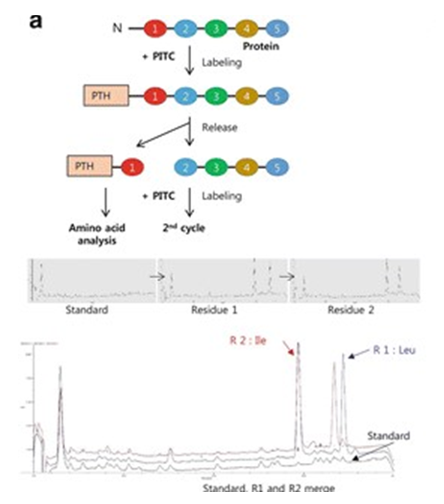
Combining [13C6]‐phenylisothiocyanate and the Edman degradation reaction: a possible breakthrough for absolute quantitative proteomics together with protein identification - Oe - 2010 - Rapid Communications in Mass Spectrometry - Wiley Online Library

Amino acid sequence assignment from single molecule peptide sequencing data using a two-stage classifier | bioRxiv
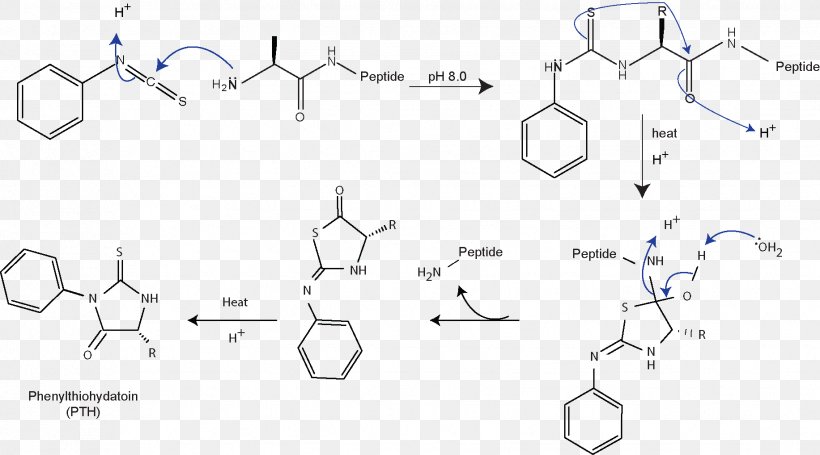
Edman Degradation Protein Sequencing Amino Acid Papain, PNG, 1833x1019px, Edman Degradation, Amino Acid, Area, Chemical Compound,

Determination of sequence and absolute configuration of peptide amino acids by HPLC–MS/CD-based detection of liberated N-terminus phenylthiohydantoin amino acids | Scientific Reports