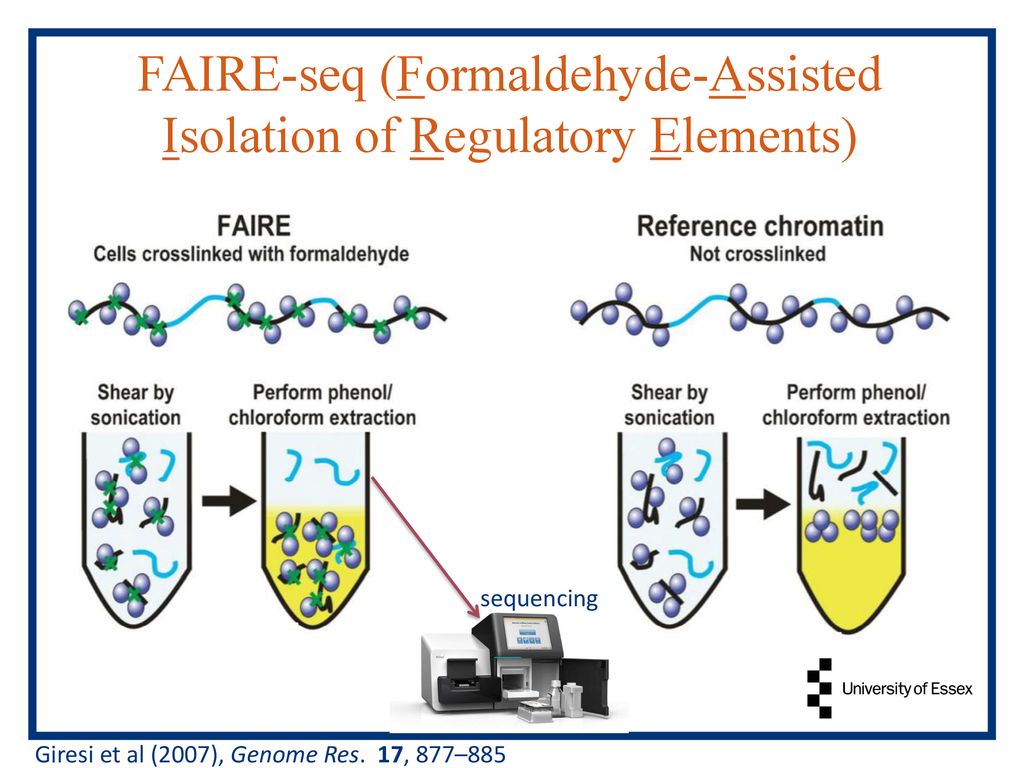
Formaldehyde-Assisted Isolation of Regulatory Elements (FAIRE) Analysis Uncovers Broad Changes in Chromatin Structure Resulting from Hexavalent Chromium Exposure | PLOS ONE

A step‐by‐step protocol for formaldehyde‐assisted isolation of regulatory elements from Arabidopsis thaliana - Omidbakhshfard - 2014 - Journal of Integrative Plant Biology - Wiley Online Library

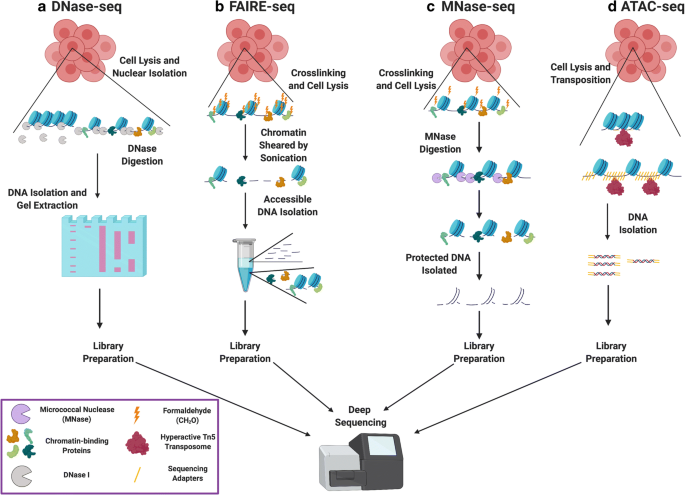
Comparing general features of ATAC-seq and FAIRE-seq. (A) Venn diagram... | Download Scientific Diagram

Example timeline for FAIRE protocol. Steps are grouped by day for the... | Download Scientific Diagram
Discovery of Transcription Factors and Regulatory Regions Driving In Vivo Tumor Development by ATAC-seq and FAIRE-seq Open Chromatin Profiling | PLOS Genetics

Isolation of active regulatory elements from eukaryotic chromatin using FAIRE (Formaldehyde Assisted Isolation of Regulatory Elements). | Semantic Scholar

Monitoring genome-wide chromatin accessibility by formaldehyde-assisted isolation of regulatory elements sequencing (FAIRE-seq) - ScienceDirect

Identifying and mitigating bias in next-generation sequencing methods for chromatin biology. - Abstract - Europe PMC