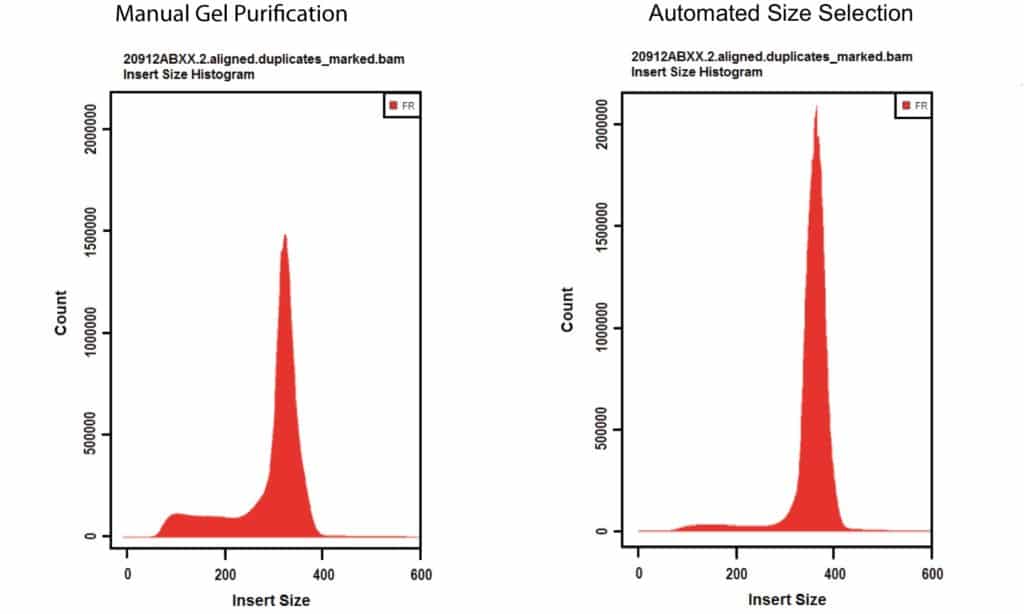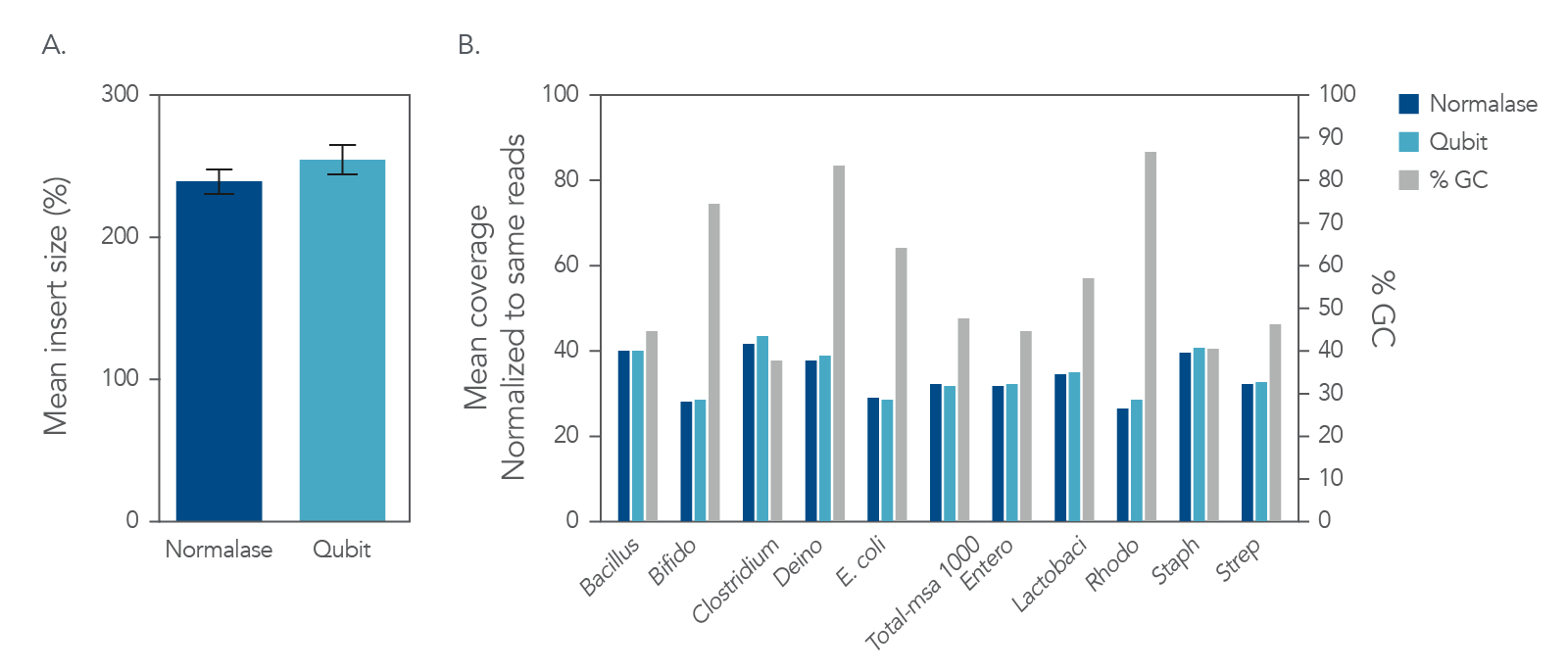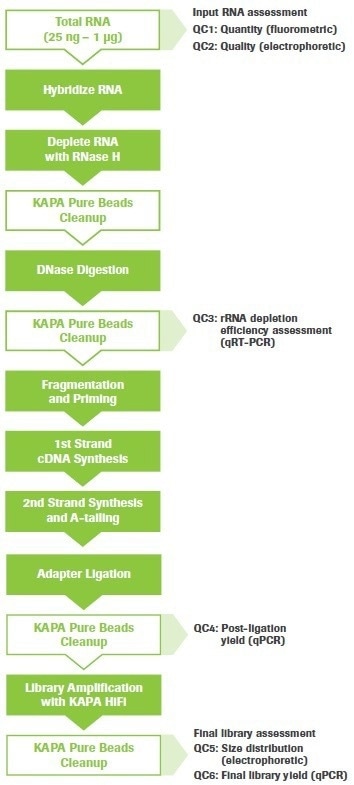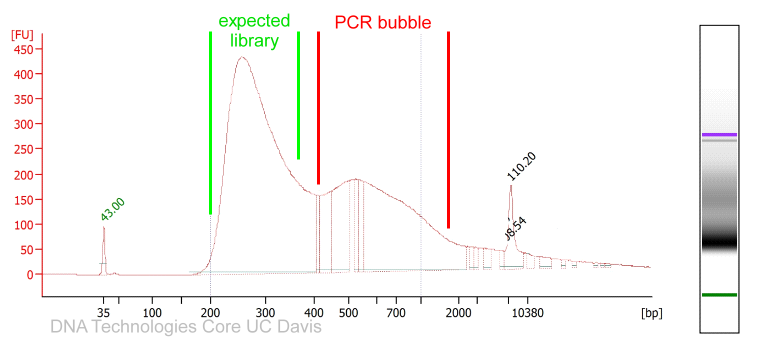
My libraries show peaks larger than expected. Can I still sequence these PCR-bubbles? | DNA Technologies Core

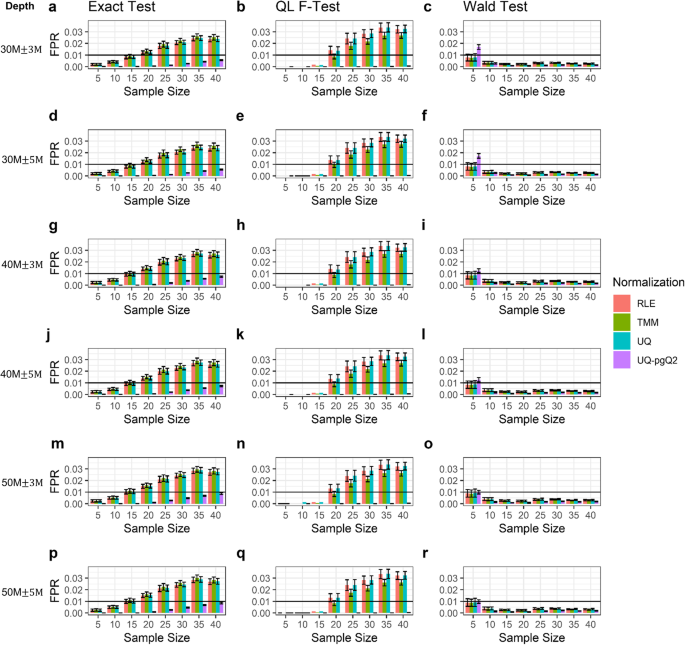
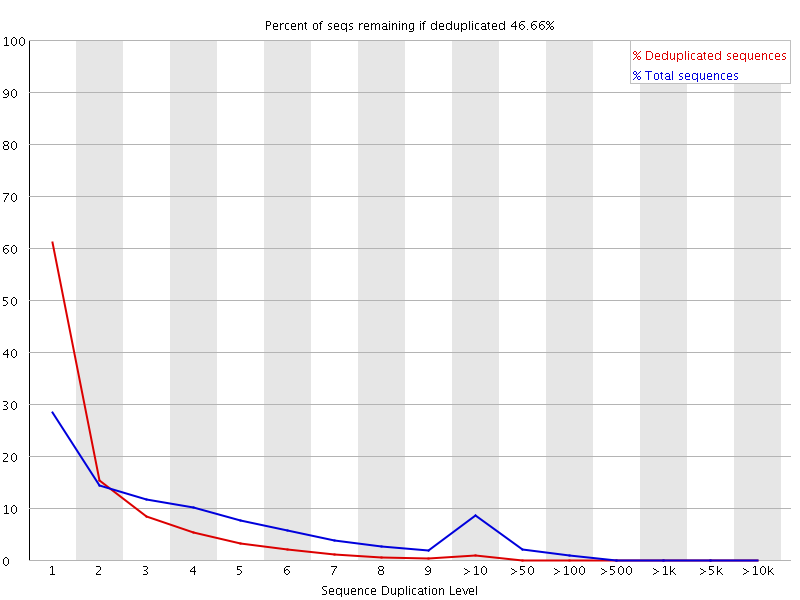
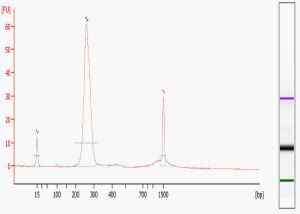
Sequencing library size distribution. Number of reads, 17 to 26 nt in... | Download Scientific Diagram
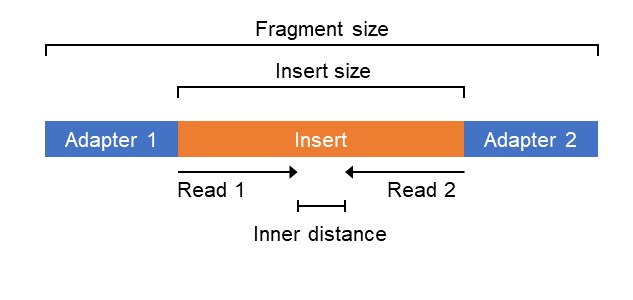
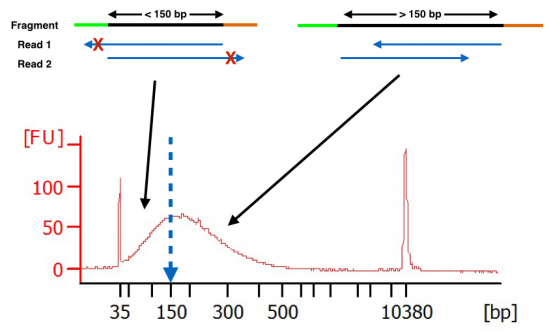
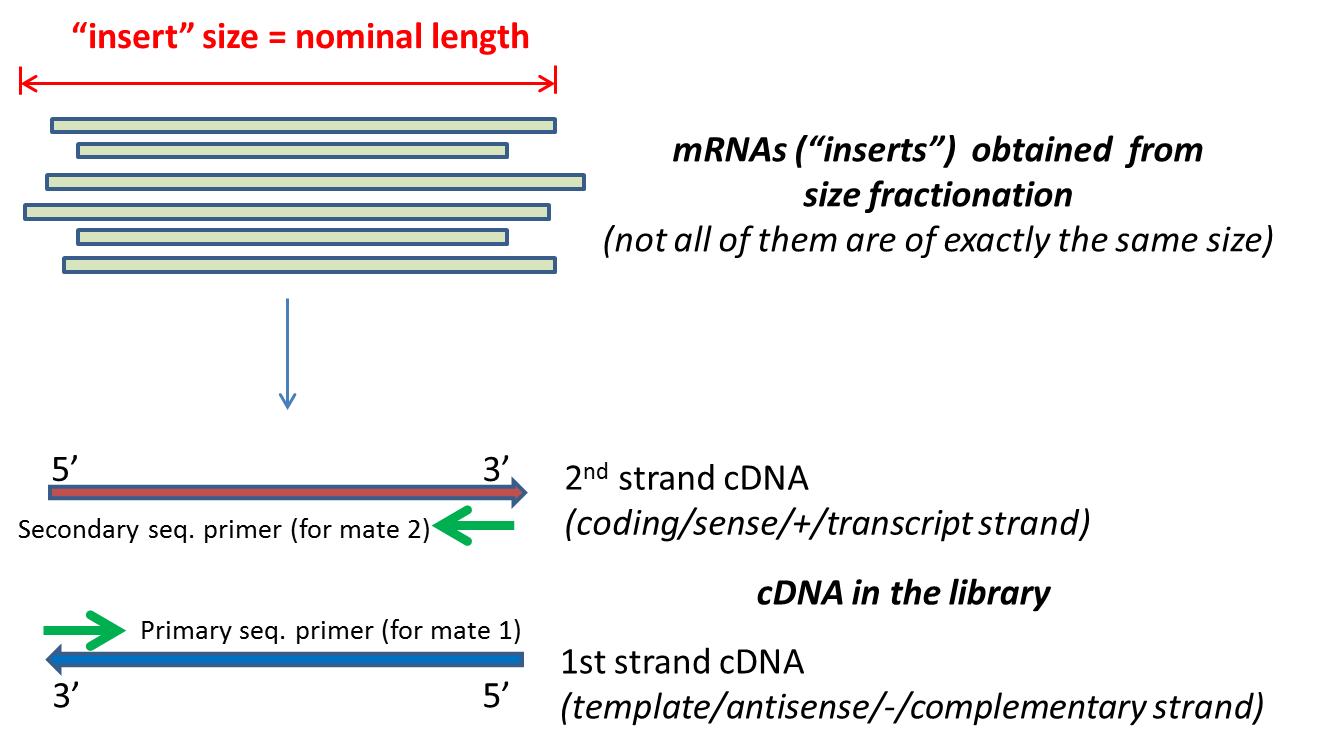
Compbio 020: Reads, fragments and inserts - what you need to know for understanding your sequencing data — Bad Grammar, Good Syntax
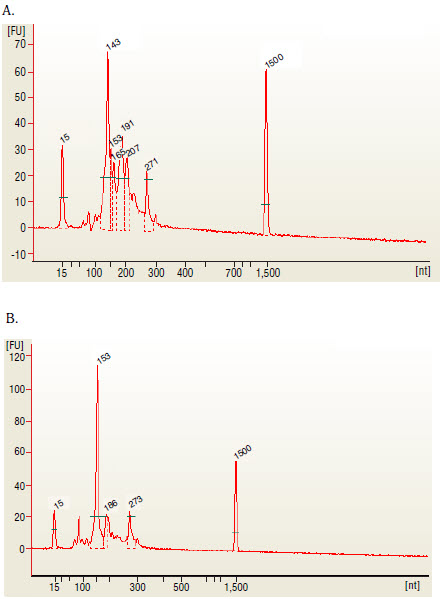
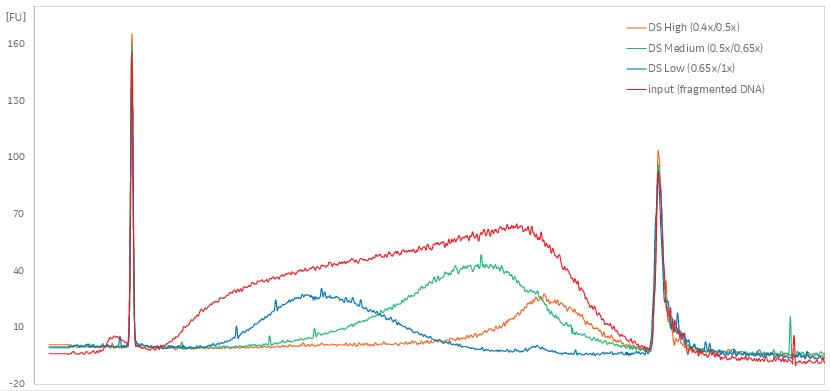
6C. Quality Control Check and Size Selection using AMPure XP Beads NEBNext Small RNA Library Prep Set for Illumina (E7300, E7580, E7560, E7330) | NEB