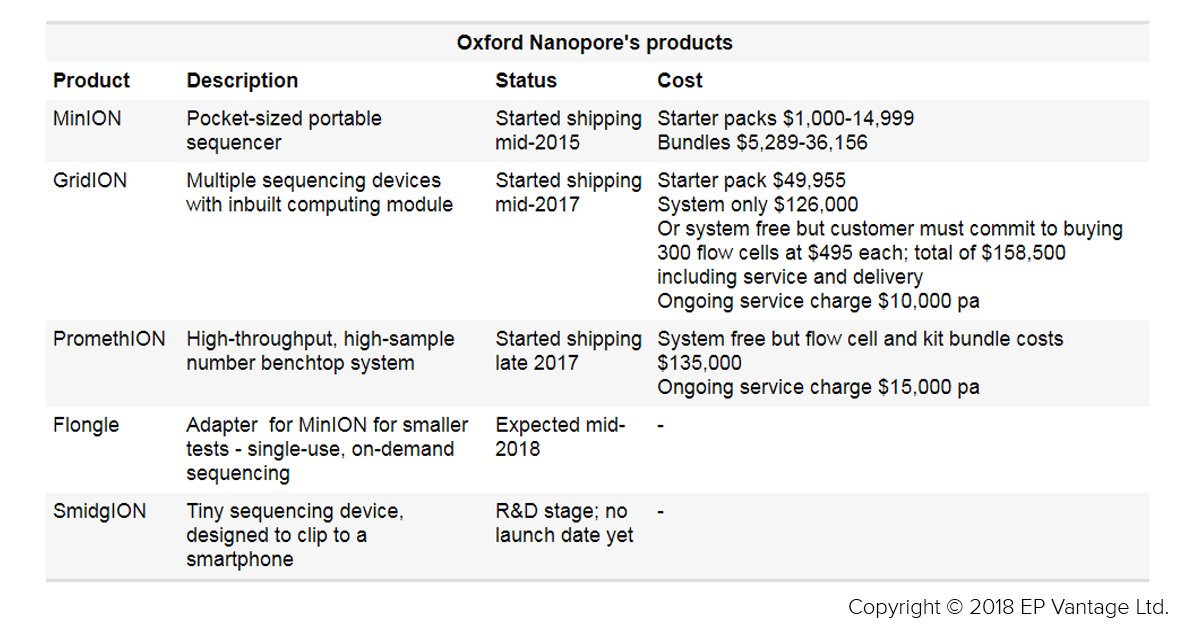
Evaluate Pharma on Twitter: "Spotlight – Oxford Nanopore, the disruptive unicorn gunning for Illumina https://t.co/azsnduXoSW via @EPVantage @LizVantage #medtech https://t.co/gory5H3qsn" / Twitter
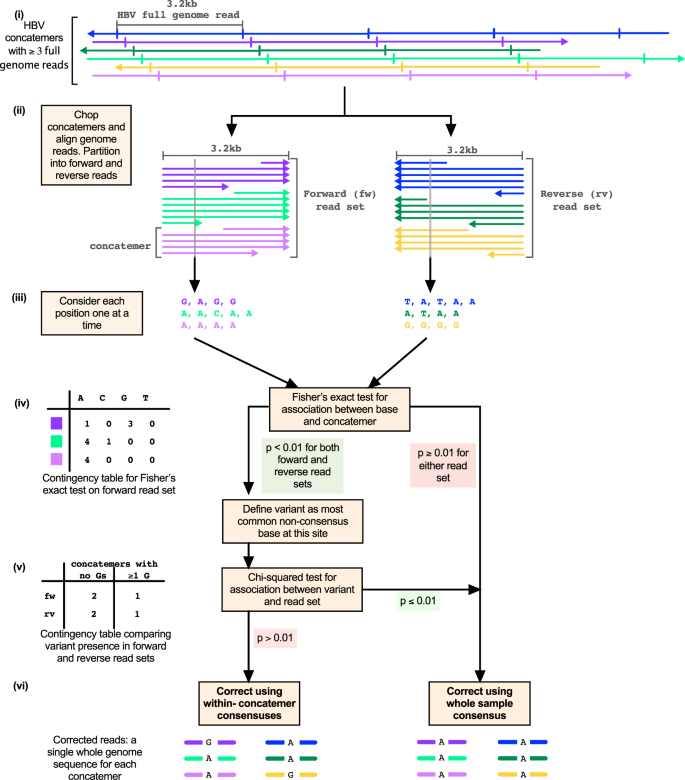
Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV) | Scientific Reports

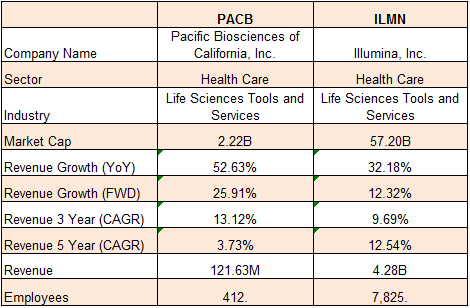
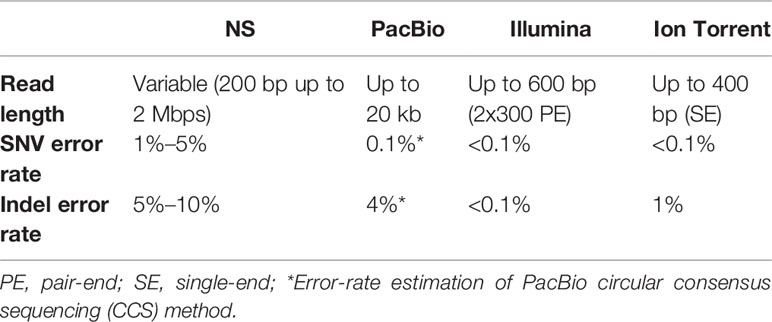
The Lady or the Tiger: Nanopore, Illumina, and PacBio – microBEnet: the microbiology of the Built Environment network

Genes | Free Full-Text | Comparison of Illumina versus Nanopore 16S rRNA Gene Sequencing of the Human Nasal Microbiota

Comparison of single nucleotide variants identified by Illumina and Oxford Nanopore technologies in the context of a potential outbreak of Shiga Toxin Producing Escherichia coli | bioRxiv

Oxford Nanopore Technology: A Promising Long-Read Sequencing Platform To Study Exon Connectivity and Characterize Isoforms of Complex Genes | Semantic Scholar

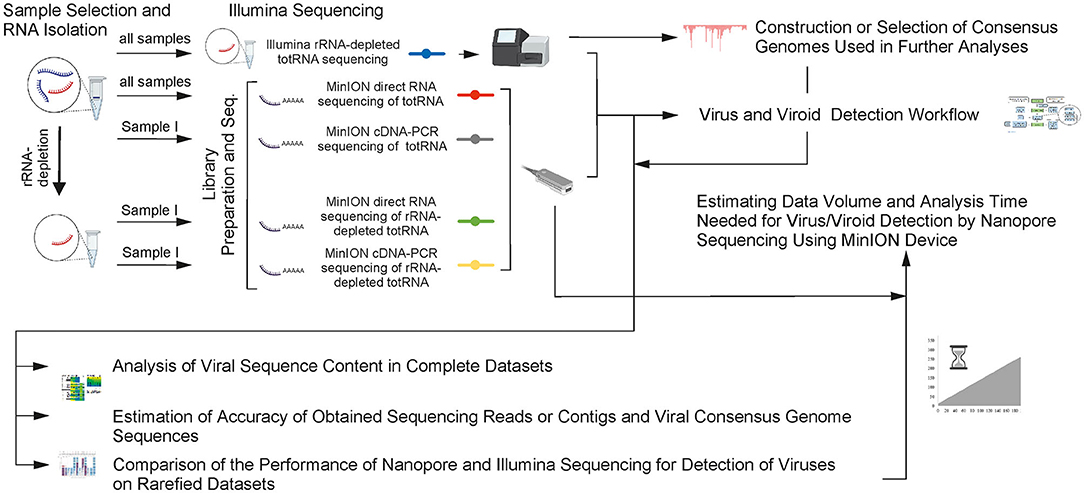
Metagenomic sequencing workflow for MinION nanopore sequencing compared... | Download Scientific Diagram

Illumina But With Nanopore: Sequencing Illumina libraries at high accuracy on the ONT MinION using R2C2 | bioRxiv
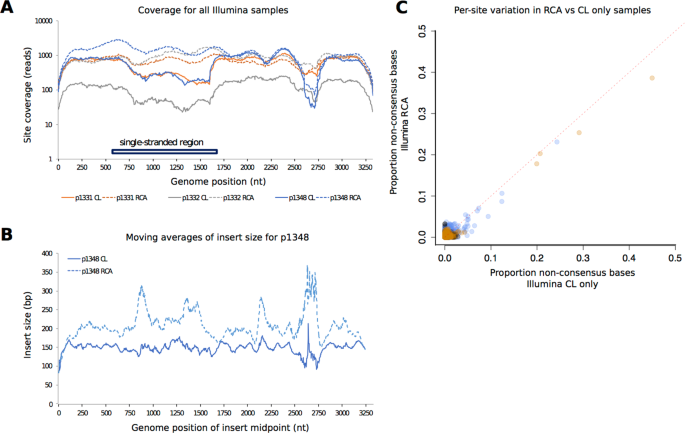
Illumina and Nanopore methods for whole genome sequencing of hepatitis B virus (HBV) | Scientific Reports