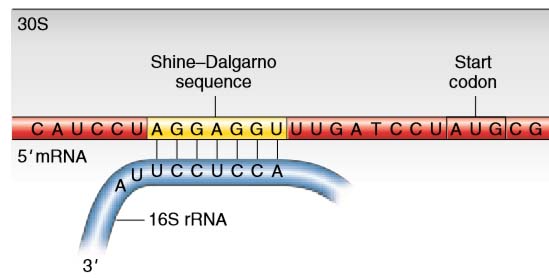
Accessibility of the Shine-Dalgarno Sequence Dictates N-Terminal Codon Bias in E. coli - ScienceDirect

Leveraging genome-wide datasets to quantify the functional role of the anti- Shine–Dalgarno sequence in regulating translation efficiency | Open Biology

An extended Shine–Dalgarno sequence in mRNA functionally bypasses a vital defect in initiator tRNA | PNAS
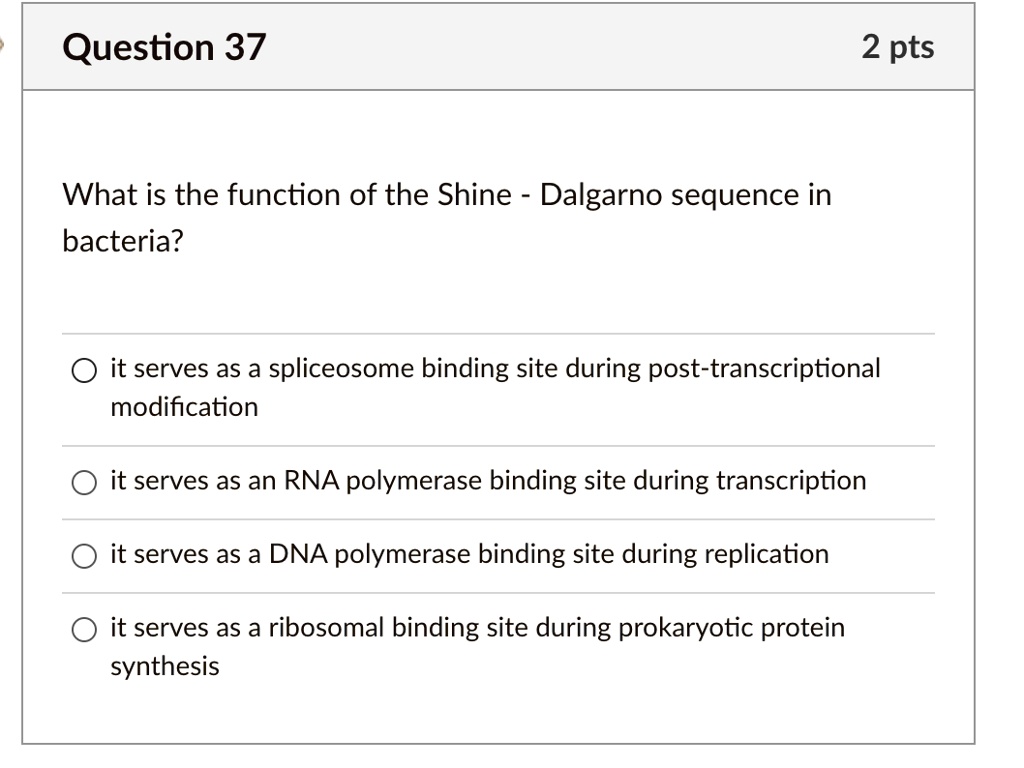

SOLVED: Question 37 2 pts What is the function of the Shine Dalgarno sequence in bacteria? it serves as a spliceosome binding site during post- transcriptional modification it serves as an RNA
Defining the anti-Shine-Dalgarno sequence interaction and quantifying its functional role in regulating translation efficiency
![PDF] Shine-Dalgarno Anti-Shine-Dalgarno Sequence Interactions and Their Functional Role in Translational Efficiency of Bacteria and Archaea | Semantic Scholar PDF] Shine-Dalgarno Anti-Shine-Dalgarno Sequence Interactions and Their Functional Role in Translational Efficiency of Bacteria and Archaea | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/ca6e3ce301334ffc5d3a7df6b8a8e5d2bc19ed16/56-Table2.1-1.png)
PDF] Shine-Dalgarno Anti-Shine-Dalgarno Sequence Interactions and Their Functional Role in Translational Efficiency of Bacteria and Archaea | Semantic Scholar
Prokaryotic coding regions have little if any specific depletion of Shine- Dalgarno motifs | PLOS ONE

The Shine-Dalgarno sequence of riboswitch-regulated single mRNAs shows ligand-dependent accessibility bursts | Nature Communications

What is the Difference Between Shine Dalgarno and Kozak Sequence | Compare the Difference Between Similar Terms
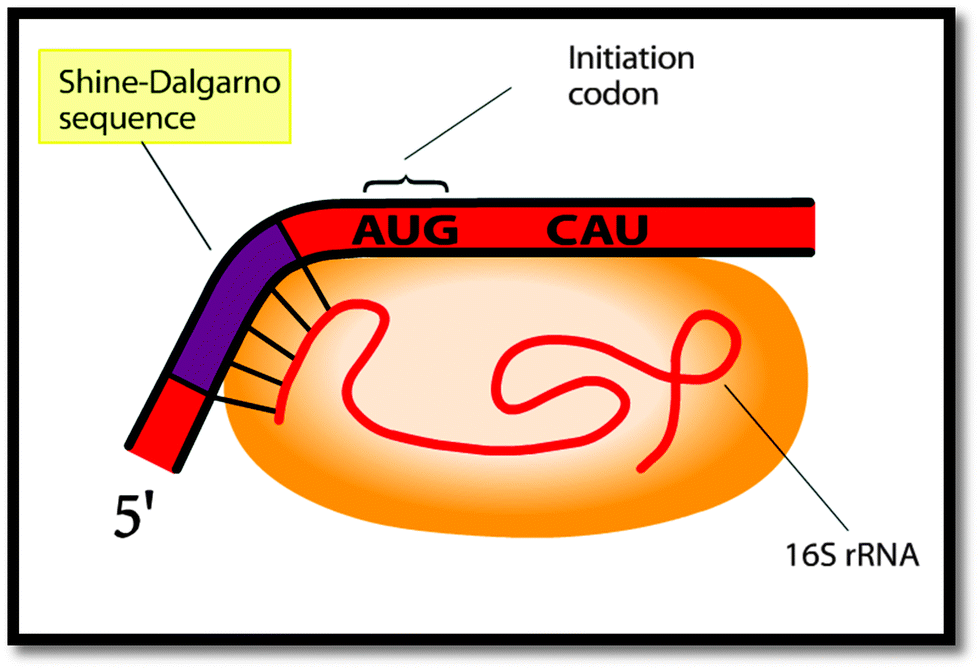
What do biochemistry students pay attention to in external representations of protein translation? The case of the Shine–Dalgarno sequence - Chemistry Education Research and Practice (RSC Publishing) DOI:10.1039/C5RP00001G
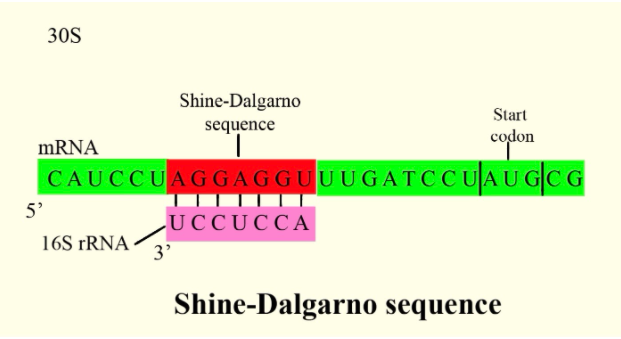
In prokaryotes, the ribosomes binding site on mRNA is called(a)Hogness sequence(b)Shine- Dalgarno sequence(c)Prinbow sequence(d)TATA- box

Defining the anti-Shine-Dalgarno sequence interaction and quantifying its functional role in regulating translation efficiency | bioRxiv