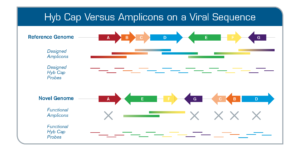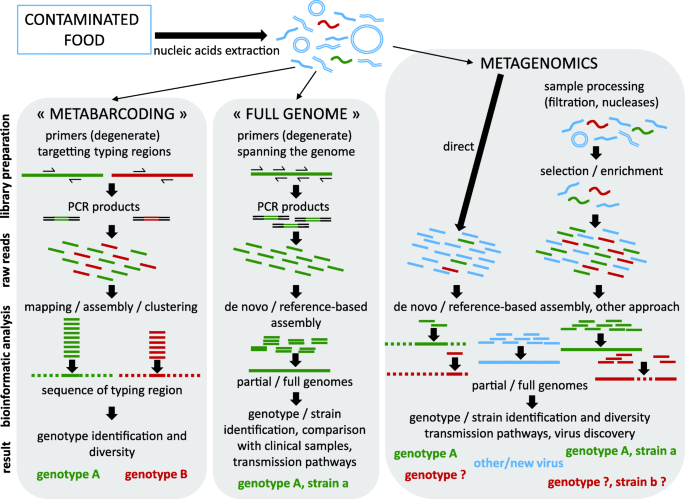
Novel opportunities for NGS-based one health surveillance of foodborne viruses | One Health Outlook | Full Text

How the Broad's Genomics Platform ramped up human and viral sequencing and kept processing COVID-19 tests, all at the same time | Broad Institute

Next-Generation Sequencing: An Eye-Opener for the Surveillance of Antiviral Resistance in Influenza: Trends in Biotechnology
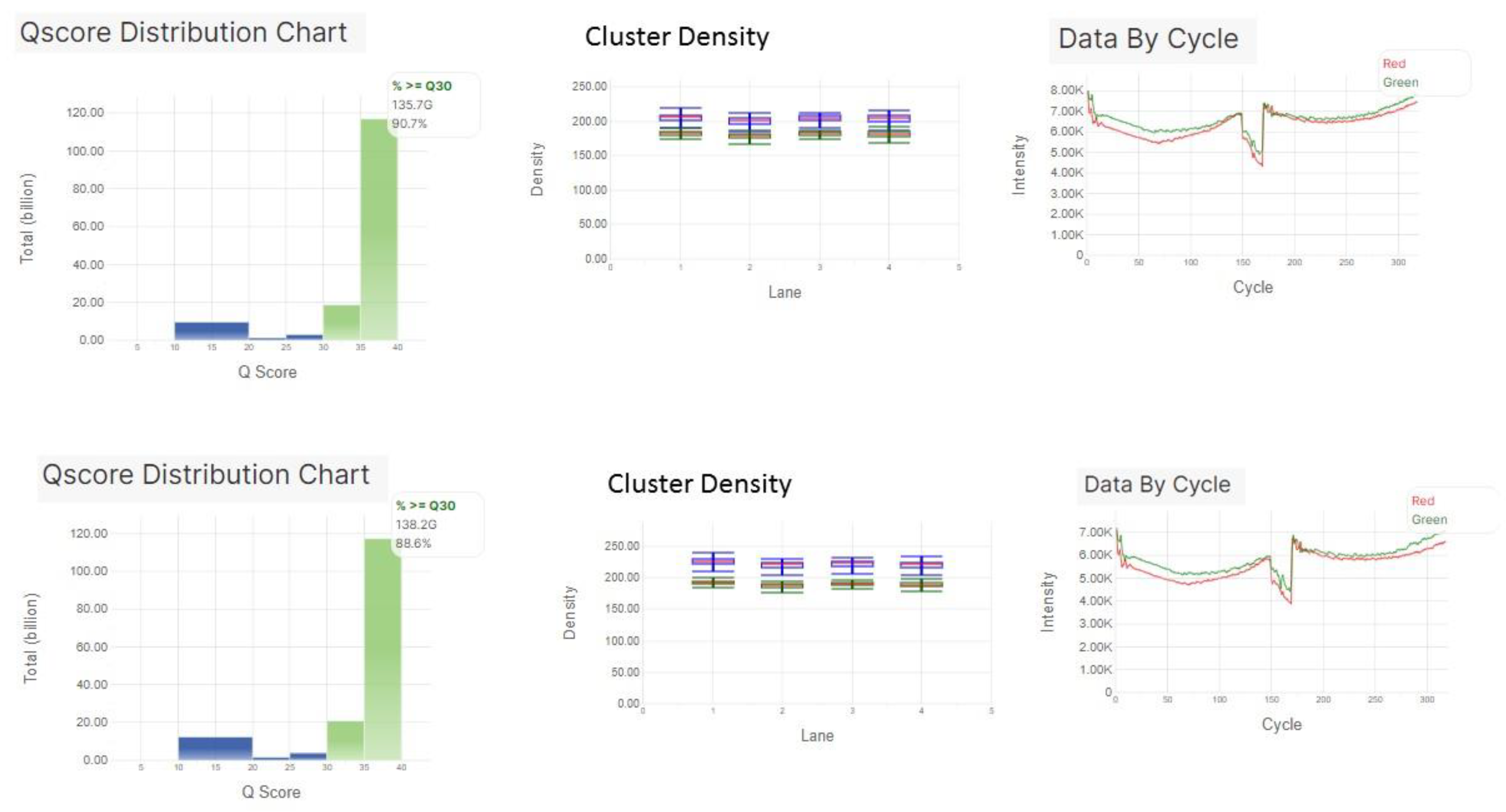
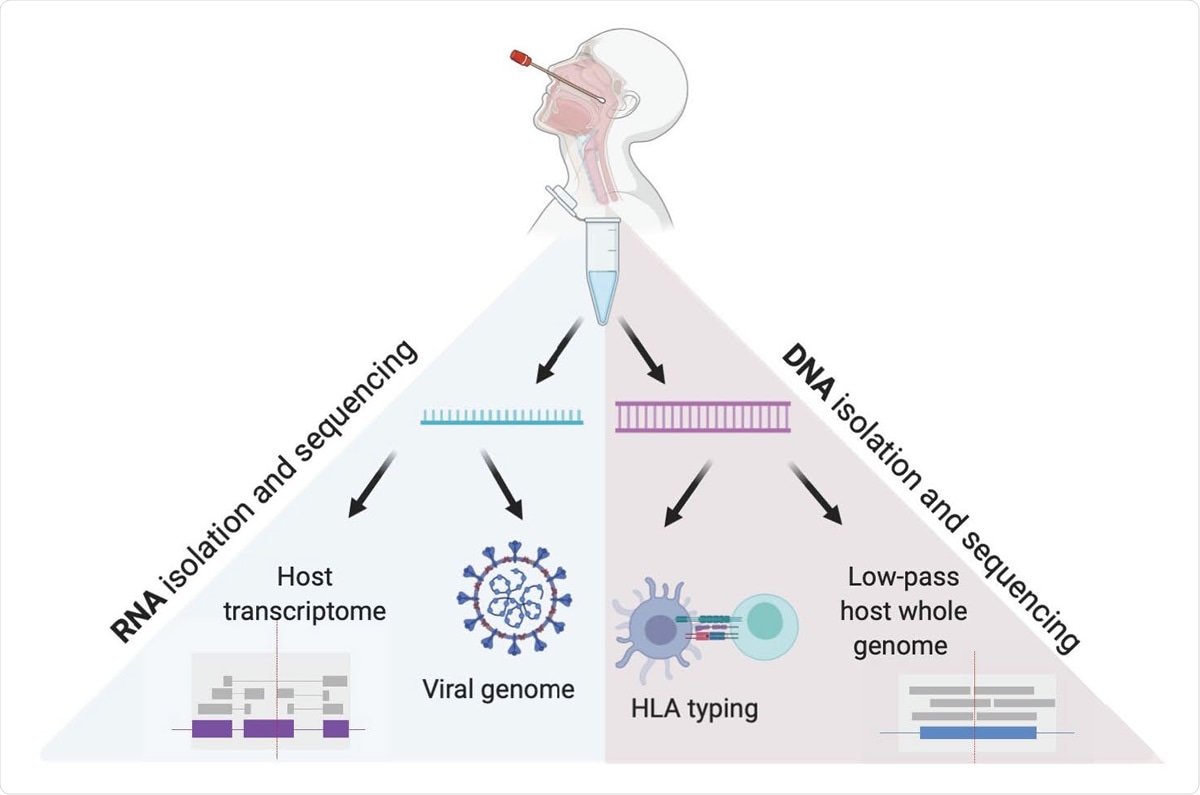
Viruses | Free Full-Text | High-Throughput Next-Generation Sequencing Respiratory Viral Panel: A Diagnostic and Epidemiologic Tool for SARS-CoV-2 and Other Viruses

Contaminating viral sequences in high-throughput sequencing viromics: a linkage study of 700 sequencing libraries - ScienceDirect

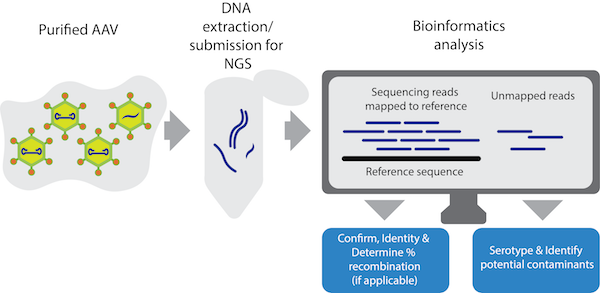
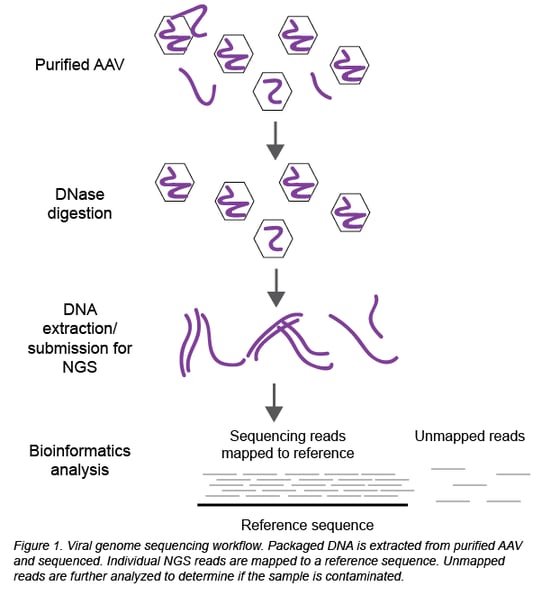
Advanced Characterization of DNA Molecules in rAAV Vector Preparations by Single-stranded Virus Next-generation Sequencing: Molecular Therapy - Nucleic Acids
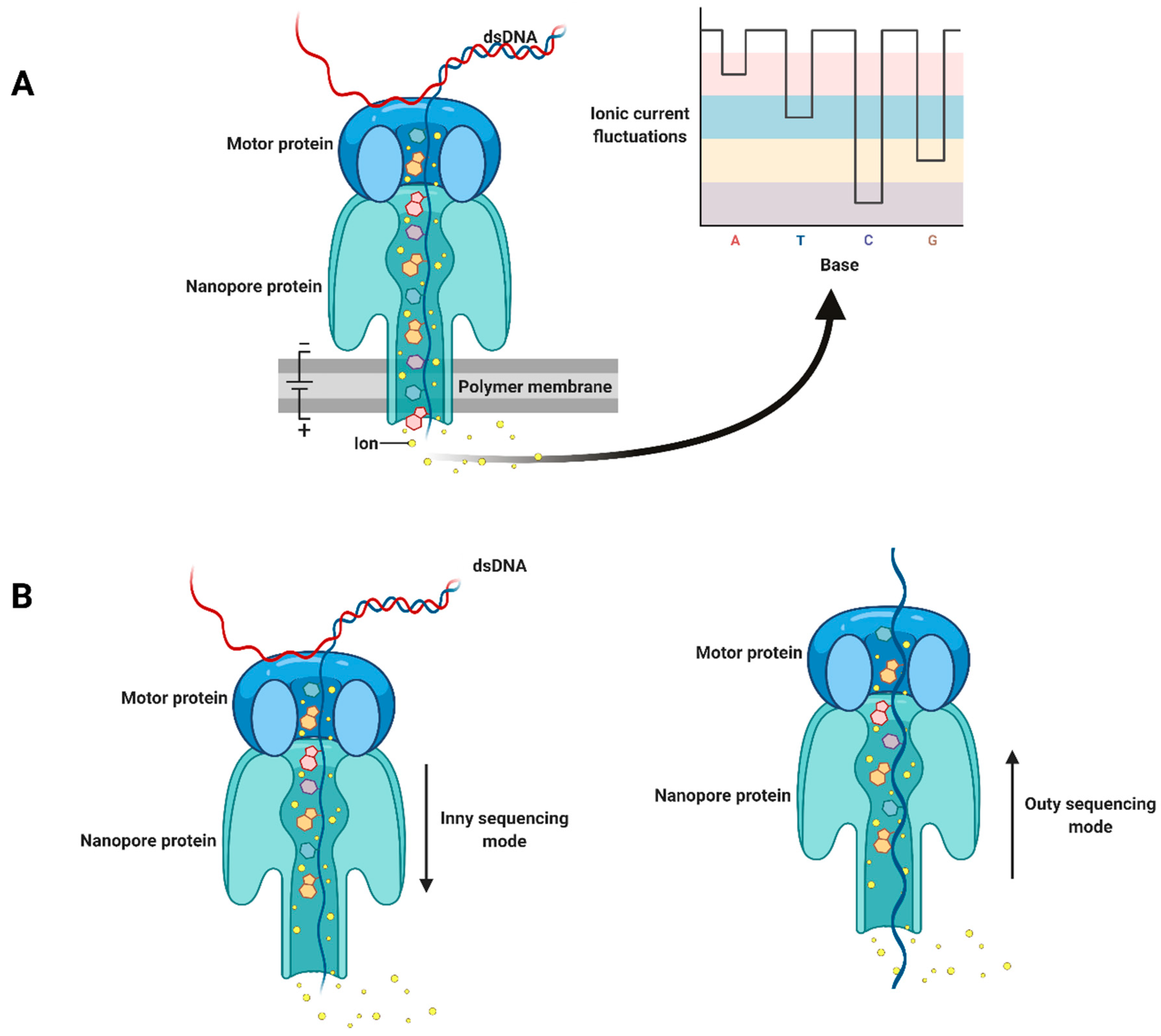
Plants | Free Full-Text | Grapevine Virology in the Third-Generation Sequencing Era: From Virus Detection to Viral Epitranscriptomics
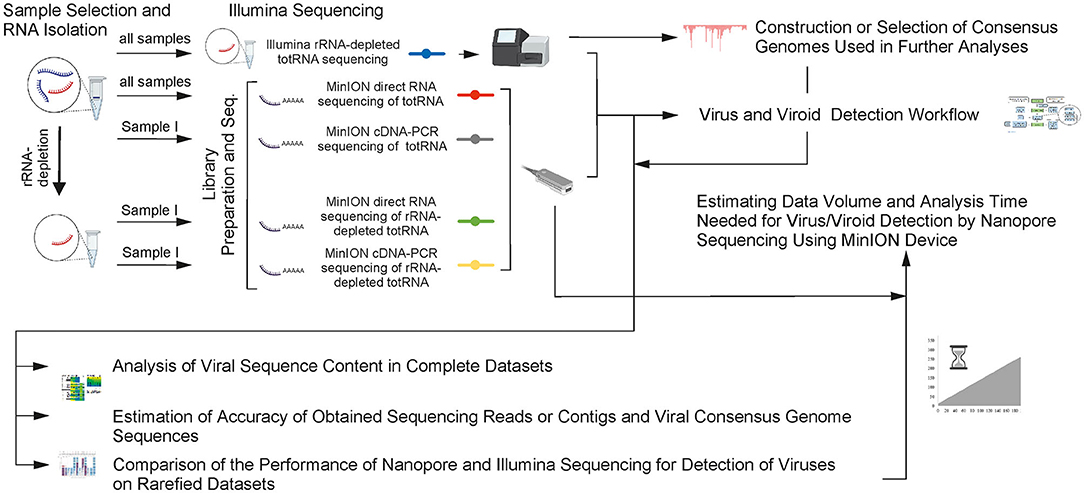
Frontiers | Systematic Comparison of Nanopore and Illumina Sequencing for the Detection of Plant Viruses and Viroids Using Total RNA Sequencing Approach

High-depth sequencing characterization of viral dynamics across tissues in fatal COVID-19 reveals compartmentalized infection | Nature Communications

Going the Distance: Optimizing RNA-Seq Strategies for Transcriptomic Analysis of Complex Viral Genomes | Journal of Virology

Tumour virology in the era of high-throughput genomics | Philosophical Transactions of the Royal Society B: Biological Sciences